Lesson 1: From Regression to Deep Learning
Learning Objectives
- Why linear models fail on complex data
- How layers + activations extend regression
- Visualize decision boundaries forming
- Forward pass intuition with minimal math
Why Regression Isn’t Enough
- Recall logistic regression decision boundaries
- Curved datasets (circles, spirals) break linear separability
- Prompt: “How could we bend this decision line?”
Notation
- Input vector: \(\mathbf{x} \in \mathbb{R}^d\)
- Weights: \(\mathbf{w} \in \mathbb{R}^d\), bias: \(b \in \mathbb{R}\)
- Activation (nonlinearity): \(\sigma(\cdot)\) e.g., ReLU, sigmoid
Tip: hidden layers typically use ReLU; outputs use sigmoid for binary classification.
Single Neuron (Scalar Output)
Linear combination + nonlinearity:
\[ \hat{y} = \sigma\big( \mathbf{w}^\top \mathbf{x} + b \big) \]
Without \(\sigma\), this is just linear regression/logistic logit.
Shapes: \(\mathbf{x}\in\mathbb{R}^d\), \(\mathbf{w}\in\mathbb{R}^d\), \(b\in\mathbb{R}\), \(\hat{y}\in\mathbb{R}\). Common \(\sigma\): ReLU, tanh, sigmoid.
Neuron + Bias Intuition
- Linear part: z = w · x + b
- Activation: a = σ(z) adds nonlinearity
- Bias b moves the decision threshold
- Layers stack these simple units
Diagram: Single Neuron
digraph neuron {
rankdir=LR;
node [shape=circle, fontsize=14, width=1.0];
edge [penwidth=1.5];
x1 [label="x₁"];
x2 [label="x₂"];
x3 [label="x₃"];
h [label="σ(w·x + b)", width=1.5];
x1 -> h;
x2 -> h;
x3 -> h;
}
Multiple inputs are combined into a single neuron that applies σ to \(w \cdot x + b\).
ReLU Activation
\(\mathrm{ReLU}(z) = \max(0, z)\) — default for hidden layers.
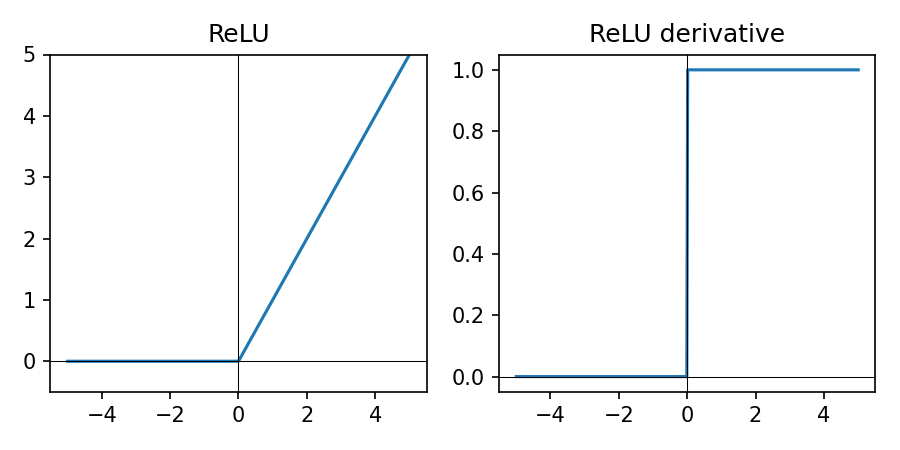
- Zero for negative inputs, linear for positive
- Keeps computation simple and fast
- Works well as a default for hidden layers
Sigmoid Activation
\(\sigma(z) = \frac{1}{1 + e^{-z}}\) — used for binary outputs.
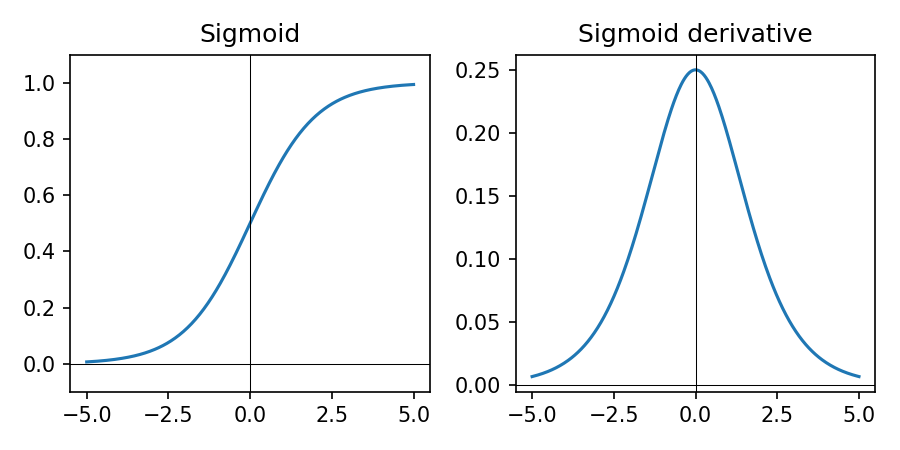
- Maps any real number to (0, 1)
- Interpretable as probability for binary class
- Used on the final neuron in this lesson
Layer (Vector Form)
Compute multiple neurons at once:
\[ \mathbf{a}^{(1)} = \sigma\big( \mathbf{W}^{(1)} \mathbf{x} + \mathbf{b}^{(1)} \big) \]
\(\mathbf{W}^{(1)} \in \mathbb{R}^{m\times d}\) maps input to \(m\) hidden units.
Shapes: \(\mathbf{b}^{(1)}\in\mathbb{R}^m\), \(\mathbf{a}^{(1)}\in\mathbb{R}^m\). \(\sigma\) applies elementwise.
Two-Layer Network
Compose layers to get flexible decision boundaries:
\[ \hat{y} = \sigma^{(2)}\!\big( \mathbf{W}^{(2)} \, \sigma^{(1)}( \mathbf{W}^{(1)}\mathbf{x} + \mathbf{b}^{(1)} ) + \mathbf{b}^{(2)} \big) \]
Each layer: linear map + nonlinearity. Stacking learns features of features.
Shapes: hidden size \(m\), output size \(k\). \(\mathbf{W}^{(2)}\in\mathbb{R}^{k\times m}\), \(\mathbf{b}^{(2)}\in\mathbb{R}^k\), \(\hat{y}\in\mathbb{R}^k\). Typical in this lesson: \(\sigma^{(1)}=\)ReLU, \(\sigma^{(2)}=\)sigmoid.
Diagram: Two-Layer Network
digraph two_layer {
rankdir=LR;
node [shape=circle, fontsize=14, width=0.8];
edge [penwidth=1.5];
graph [nodesep=0.5, ranksep=1.0];
subgraph cluster_input {
label="Inputs";
color="white";
x1 [label="x₁"];
x2 [label="x₂"];
}
subgraph cluster_hidden {
label="Hidden layer (ReLU)";
style=filled; fillcolor="#f5f5f5";
h1 [label="h₁"];
h2 [label="h₂"];
h3 [label="h₃"];
}
subgraph cluster_output {
label="Output";
color="white";
y [label="ŷ"];
}
x1 -> h1; x1 -> h2; x1 -> h3;
x2 -> h1; x2 -> h2; x2 -> h3;
h1 -> y; h2 -> y; h3 -> y;
}
Inputs feed into hidden units, which then feed into the output neuron.
Forward Pass — Step by Step
- Compute each layer’s z = W·x + b
- Apply activation a = σ(z)
- Feed a into next layer
- Final output → prediction
We’ll worry about learning (backprop) next lesson.
Key Themes
- Linear part learns weighted combinations; bias shifts thresholds
- Nonlinearity lets boundaries bend (beyond any single line)
- Depth composes simple units into complex patterns
Interactive Demo (TF Playground)
- Select circular or spiral dataset
- No hidden layers → poor separation
- 1 hidden layer (8) → improvement
- 2 layers (8+8) → complex shapes classified
- Tune activations, learning rate, noise
Discuss: Why deeper → better patterns? What might each neuron learn?
Mini Hands-On
Implement the idea in PyTorch on synthetic 2D points.
import torch
import torch.nn as nn
model = nn.Sequential(
nn.Linear(2, 8),
nn.ReLU(),
nn.Linear(8, 8),
nn.ReLU(),
nn.Linear(1, 1),
nn.Sigmoid(),
)
# optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
# loss_fn = nn.BCELoss()
# training loop ...
Plot decision regions; relate to Playground behavior.
Code: Make a 2D Dataset
# Option A: use scikit-learn (concise)
from sklearn.datasets import make_circles
import numpy as np
X, y = make_circles(n_samples=1000, factor=0.5, noise=0.1, random_state=0)
X = X.astype('float32')
y = y.astype('float32')
# Option B: quick NumPy spiral (for the curious)
def make_spiral(n=500, noise=0.2):
n2 = n//2
t = np.linspace(0, 2*np.pi, n2)
r = np.linspace(0.2, 1.0, n2)
x1 = np.c_[r*np.cos(t), r*np.sin(t)] + noise*np.random.randn(n2,2)
x2 = np.c_[-r*np.cos(t), -r*np.sin(t)] + noise*np.random.randn(n2,2)
Xs = np.vstack([x1, x2]).astype('float32')
ys = np.r_[np.zeros(n2), np.ones(n2)].astype('float32')
return Xs, ys
Code: Train and Evaluate (PyTorch)
import torch
import torch.nn as nn
X_t = torch.from_numpy(X)
y_t = torch.from_numpy(y).unsqueeze(1)
model = nn.Sequential(
nn.Linear(2, 8),
nn.ReLU(),
nn.Linear(8, 8),
nn.ReLU(),
nn.Linear(8, 1),
nn.Sigmoid(),
)
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
loss_fn = nn.BCELoss()
for epoch in range(20):
optimizer.zero_grad()
preds = model(X_t)
loss = loss_fn(preds, y_t)
loss.backward()
optimizer.step()
print('Final training loss:', float(loss))
Expect validation accuracy to improve over epochs; details next lesson.
Code: Plot Decision Regions
import numpy as np, matplotlib.pyplot as plt
import torch
xx, yy = np.meshgrid(np.linspace(X[:,0].min()-0.5, X[:,0].max()+0.5, 200),
np.linspace(X[:,1].min()-0.5, X[:,1].max()+0.5, 200))
grid = np.c_[xx.ravel(), yy.ravel()].astype('float32')
with torch.no_grad():
grid_t = torch.from_numpy(grid)
probs = model(grid_t).detach().numpy().reshape(xx.shape)
plt.figure(figsize=(5,4))
plt.contourf(xx, yy, probs, levels=20, cmap='RdBu', alpha=0.6)
plt.scatter(X[:,0], X[:,1], c=y, cmap='RdBu', edgecolor='k', s=12)
plt.title('Decision regions')
plt.show()
Relate the learned boundary to the Playground visuals.
Wrap-Up
- Neuron = regression + nonlinearity on top
- Layers stack neurons to learn features of features
- Depth + activations bend decision boundaries into complex shapes
Homework:
- In the spirals notebook, recreate one of the Playground experiments (spirals or circles).
- Try at least two different architectures.
- Find the deepest network (most layers) that uses the fewest units but still fits the data.
- Find the shallowest network (fewest layers) that uses the fewest units but still fits the data.
Next time: how networks actually learn these weights using loss functions and gradient descent.